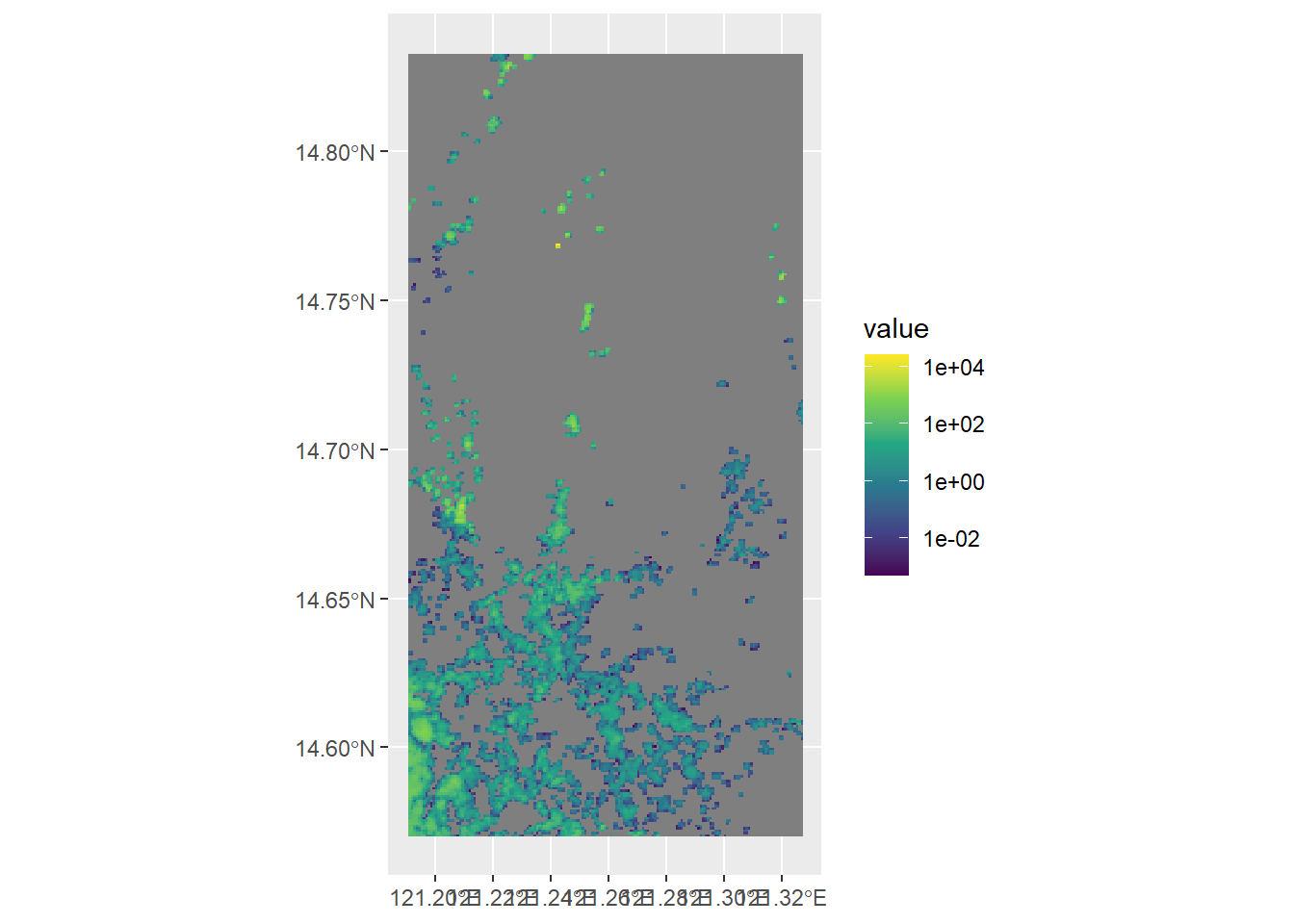
# download file from GHS database
if (!dir.exists("data/ghsl")) dir.create("data/ghsl")
file <- "GHS_POP_E2020_GLOBE_R2023A_4326_3ss_V1_0_R8_C31"
paste0(
"https://jeodpp.jrc.ec.europa.eu/ftp/jrc-opendata/GHSL/",
"GHS_POP_GLOBE_R2023A/GHS_POP_E2020_GLOBE_R2023A_4326_3ss/V1-0/tiles/",
file, ".zip"
) |>
download.file(paste0("data/ghsl/", file, ".zip"), method = "curl")
unzip(paste0("data/ghsl/", file, ".zip"), exdir = "data/ghsl")
# crop map
area <- list.files("data/umrbpl/", full.names = TRUE, pattern = "shp$") |>
read_sf() |>
st_transform(crs = "EPSG:4326")
pop <- paste0("data/ghsl/", file, ".tif") |>
rast() |>
crop(area)
writeRaster(pop, file = "data/ghsl/population_umrbpl.tif")